Imaris Open
FILE EXCHANGE & FORUM
Frequently Asked Questions
Imaris 9.8 brings several usability improvements for its users, including easier volume visualisation modes and virtual sample sectioning (clipping planes), which are particularly applicable when visualizing large and thick samples, like those acquired with light sheet microscopy. At the same time this release brings new possibilities for object detection validation and editing in dense datasets as object boundaries are rendered together with the microscopy signal on a slicer. Neuroscientists analysing neurons or microglia will appreciate the fact that the popular Filament Tracer module was enriched with a soma model which ensures optimal reconstruction and measurements of somas and dendrites.
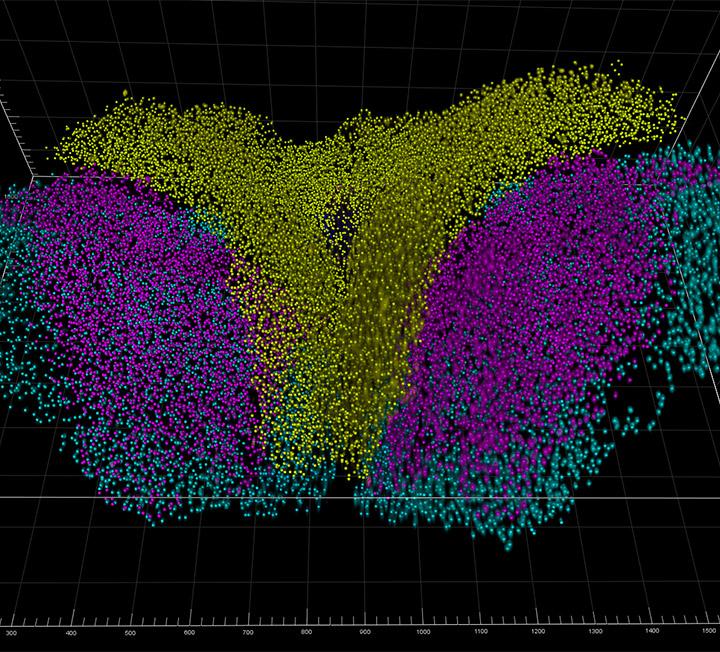
Try Imaris 9.8 for FreeSetting up the segmentation parameters in crowded datasets is no longer a challenging task in Imaris 9.8. This release brings new functionality to render objects on slicers, including Spots, Surfaces, Filaments and Tracks, which is essential in thick datasets with multiple objects, such as light sheet images, OPT, thick confocal sections or multiphoton datasets.
3D View
2D Slicer View
3D View
2D Slicer View
3D View

2D Slicer View
In visualizing microscopy data it’s equally important to see the 3D rendering as to make the virtual dissection of the sample to see what’s inside the structure.
Imaris 9.8 introduces a new handling method for clipping planes and slicers to improve efficiency and enable better visualization flow for Imaris users. The new functionality improves visualization options for large 3D data volumes. Both clipping plane and slicers are now positioned using a very precise and intuitive 3-arrow manipulator. In Imaris users can visualize microscopy datasets in the terabyte range (3.5 TB data visualization tested in house; up to 20 TB visualization reported by the users) as well as in gigabytes range.
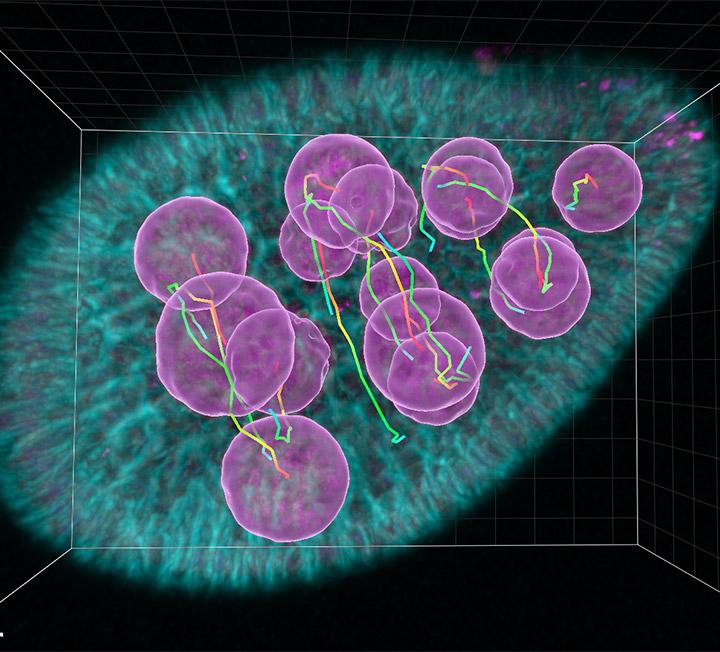
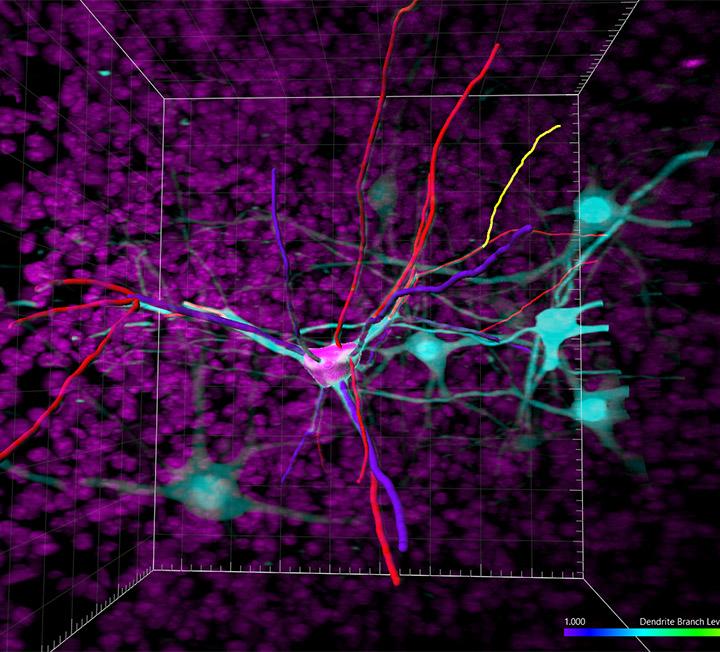
For accurate tracing of dendrites and microglia processes, one needs to model cell bodies – somas.
Imaris 9.8 improves its widely known Filament Tracer module with effective soma detection to bring the most accurate dendrite and soma measurements to life. In addition, the speed of rendering filaments is much improved making interaction with your images smoother and easier.
The complete neuron model: soma, axon, dendrites and dendritic spines can be rendered in 3D and on the extended 2D section. Rendering on the 2D section is extremely useful when tracing dense neuronal structures and to validate object detection.

After having used Imaris 9.8 for just 5 minutes, I cannot even remember anymore how I used Imaris before that. Thus, an absolutely great addition to the software!
Christian Tischer, PhD, EMBL’s Centre for Bioimage Analysis (CBA), Heidelberg, Germany
The Imaris Learning Center hosts a wide range of tutorial videos, how-to articles and webinars to guide you through the many features of Imaris. We have provided some links below which will get you started on some of our most recent developments.